Structurally similar motifs in the (H2A-H2B) and the (H3-H4) histone... | Download Scientific Diagram
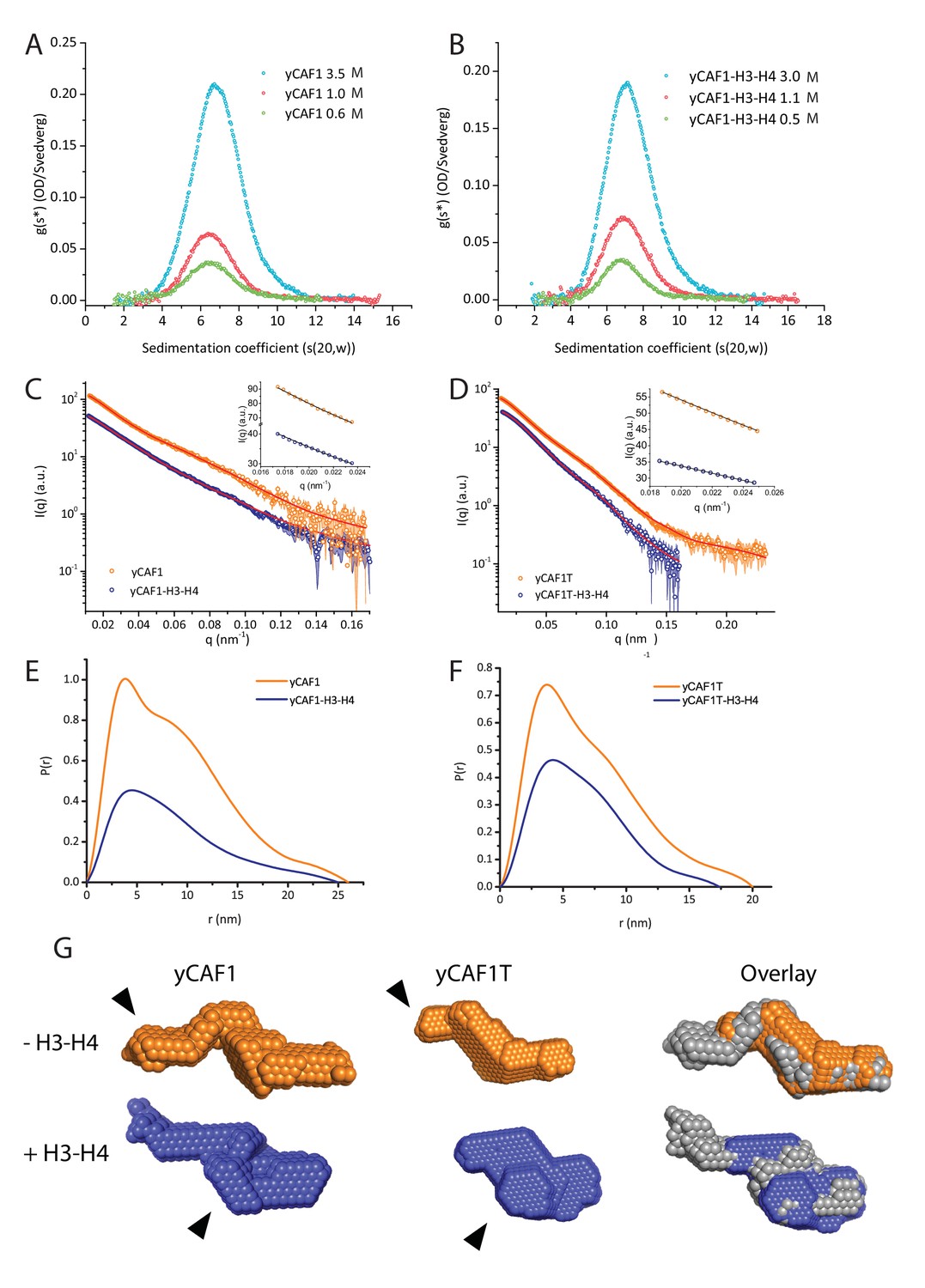
Insights into the molecular architecture and histone H3-H4 deposition mechanism of yeast Chromatin assembly factor 1 | eLife

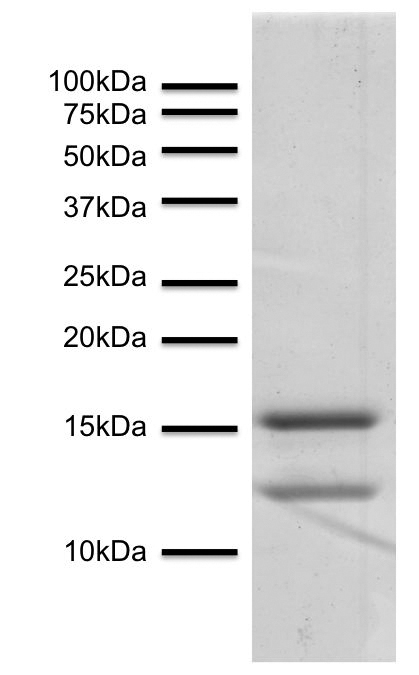
Simultaneous Proteoform Analysis of Histones H3 and H4 with a Simplified Middle-Down Proteomics Method | Analytical Chemistry

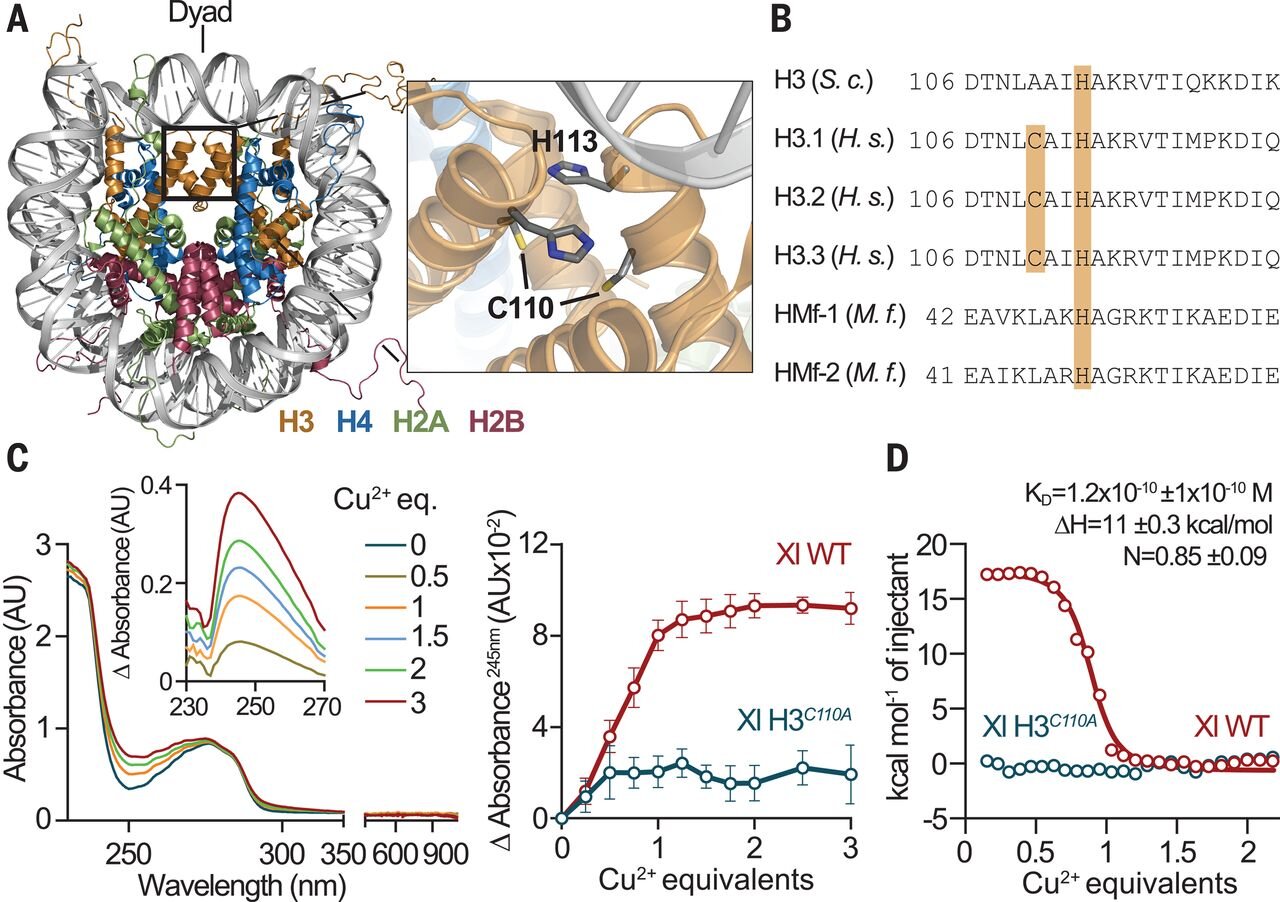
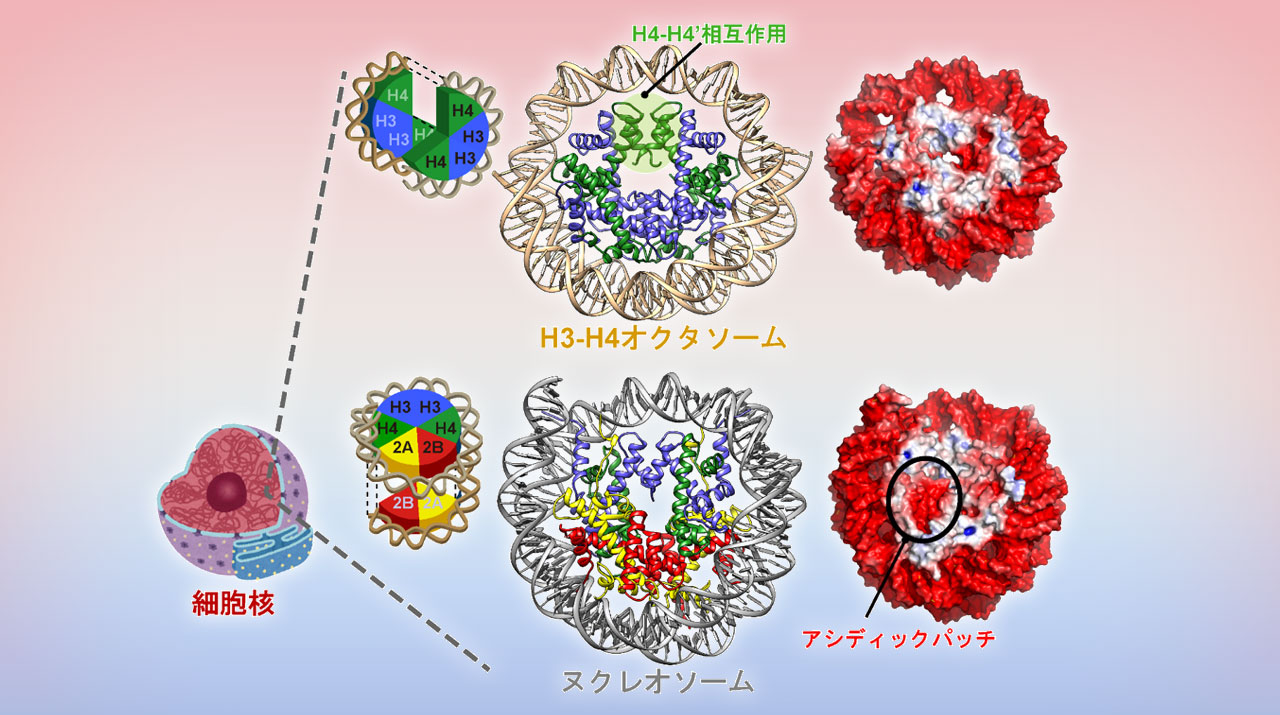
Cryo–electron microscopy structure of the H3-H4 octasome: A nucleosome-like particle without histones H2A and H2B | PNAS
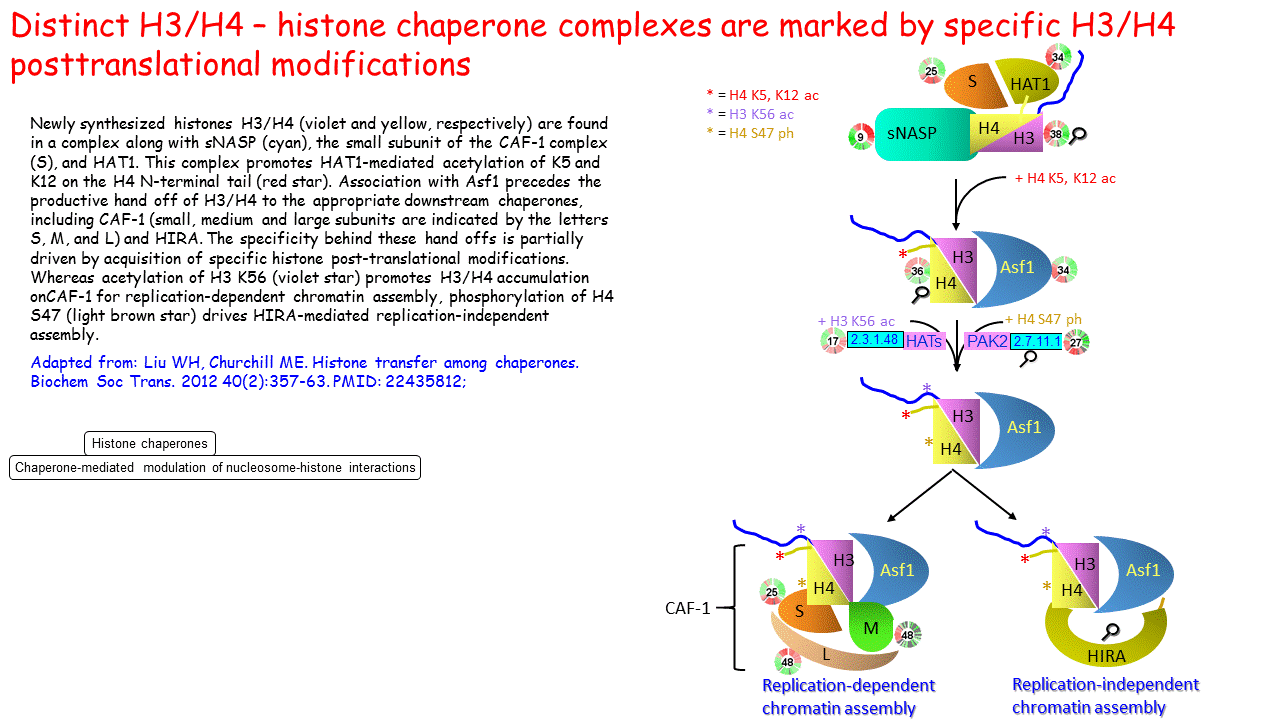
Distinct H3/H4 – histone chaperone complexes are marked by specific H3/H4 posttranslational modifications

Cryo–electron microscopy structure of the H3-H4 octasome: A nucleosome-like particle without histones H2A and H2B | PNAS

POLE3-POLE4 Is a Histone H3-H4 Chaperone that Maintains Chromatin Integrity during DNA Replication - ScienceDirect

Schematic Representation of DNA Wound Around a Central Histone H3/H4... | Download Scientific Diagram

Distinct H3/H4 – histone chaperone complexes are marked by specific... | Download Scientific Diagram

Insights into the molecular architecture and histone H3-H4 deposition mechanism of yeast Chromatin assembly factor 1 | eLife

MCM2 binding to histones H3–H4 and ASF1 supports a tetramer-to-dimer model for histone inheritance at the replication fork | Nature Structural & Molecular Biology